Using single cell technology, a new study sheds light on the significance of genetic risk factors for, and the diversity of cells involved in, the development of coronary artery disease. The researchers analysed human atherosclerotic lesions to map the chromatin accessibility of more than 7,000 cells. The chromatin accessibility is known to reflect active regions and genes in the genome. The findings were published in Circulation Research.
Genome-wide association studies of the human genome have identified over 200 loci associated with coronary artery disease. More than 90% of them are located outside protein-coding genes, in so called cis-regulatory elements, whose significance in the pathogenesis of coronary artery disease remains unclear.
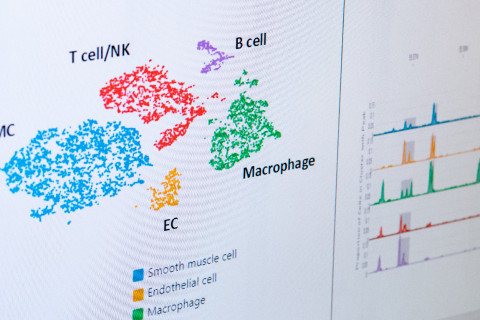
Previous research has demonstrated that the development of coronary artery disease involves a variety of different cells and their subtypes. The now-published study is the first to use single cell technology to map epigenetic changes in these cells. The researchers used the ATAC-seq sequencing method to discover the nuclear chromatin structure of endothelial cells and smooth muscle cells, as well as immune system monocytes, macrophages, NK/T and B cells, providing a unique resource to study the cell-type specific activity of the cis-regulatory elements in the disease affected vessel wall.
The study demonstrated that genetic risk variants associated with coronary artery disease are particularly enriched in cis-regulatory elements specific to endothelial and smooth muscle cells, indicating that these cells play a significant role in transmitting susceptibility to the disease. Based on chromatin accessibility mapping and gene expression data, the researchers were able to identify putative target genes for approximately 30% of all known loci associated with coronary artery disease. In addition, the researchers performed genome-wide experimental fine-mapping of the variants, allowing them to identify potential causal single-nucleotide polymorphisms and the associated target gene for over 30 loci that have been linked to coronary artery disease. The study also presented a number of examples of how the chromatin accessibility and gene expression data can be used to predict target cells via which the function of the genetic changes associated with the disease is transmitted in the tissue. This is a significant step forward that helps to understand the real functional significance of risk variants in the pathophysiology of coronary artery disease. In the future, this information can be used to develop more effective, safer and more individualised treatments for coronary artery disease.
The study was conducted in the Cardiovascular Genomics laboratory led by Associate Professor Minna Kaikkonen-Määttä at the University of Eastern Finland, in close collaboration with the research groups of Tapio Lönnberg (University of Turku), Seppo Ylä-Herttuala (University of Eastern Finland), Mete Civelek (University of Virginia), and Casey Romanoski (University of Arizona).The study was funded by the European Research Council (ERC) under the European Union’s Horizon 2020 research and innovation programme (Grant No. 802825), the Academy of Finland, the Sigrid Jusélius Foundation, the American Heart Association, and the US National Institutes of Health.
Research article:
Örd T, Õunap K, Stolze L, Aherrahrou R, Nurminen V, Selvarajan I, Toropainen A, Lönnberg T, Aavik E, Ylä-Herttuala S, Civelek M, Romanoski CE, Kaikkonen MU. Single-Cell Epigenomics and Functional Fine-Mapping of Atherosclerosis GWAS Loci. Circ Res. 2021 May 24. doi: 10.1161/CIRCRESAHA.121.318971. Epub ahead of print. PMID: 34024118.
https://pubmed.ncbi.nlm.nih.gov/34024118/